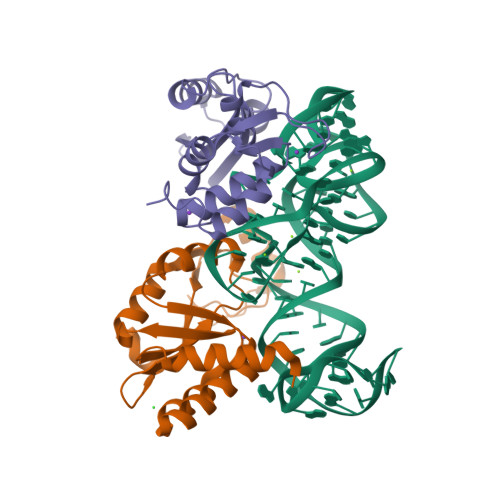
RCSB PDB - 5DAR: CRYSTAL STRUCTURE OF THE BASE OF THE RIBOSOMAL P STALK FROM METHANOCOCCUS JANNASCHII

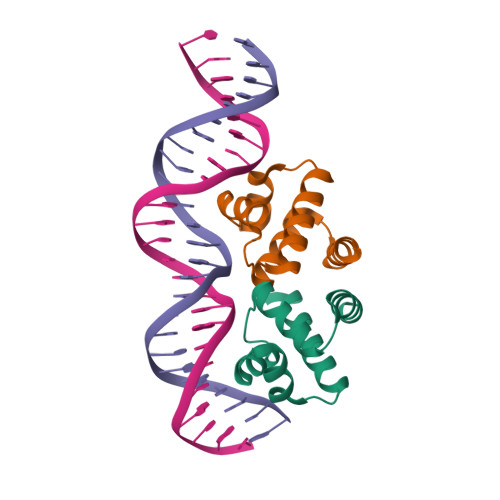
RCSB PDB - 1LCD: STRUCTURE OF THE COMPLEX OF LAC REPRESSOR HEADPIECE AND AN 11 BASE-PAIR HALF-OPERATOR DETERMINED BY NUCLEAR MAGNETIC RESONANCE SPECTROSCOPY AND RESTRAINED MOLECULAR DYNAMICS

RCSB PDB - 1FUF: CRYSTAL STRUCTURE OF A 14BP RNA OLIGONUCLEOTIDE CONTAINING DOUBLE UU BULGES: A NOVEL INTRAMOLECULAR U*(AU) BASE TRIPLE

RCSB PDB - 1GQU: Crystal structure of an alternating A-T oligonucleotide fragment with Hoogsteen base pairing

RCSB PDB - 2BR0: DNA Adduct Bypass Polymerization by Sulfolobus solfataricus Dpo4. Analysis and Crystal Structures of Multiple Base-Pair Substitution and Frameshift Products with the Adduct 1,N2-Ethenoguanine

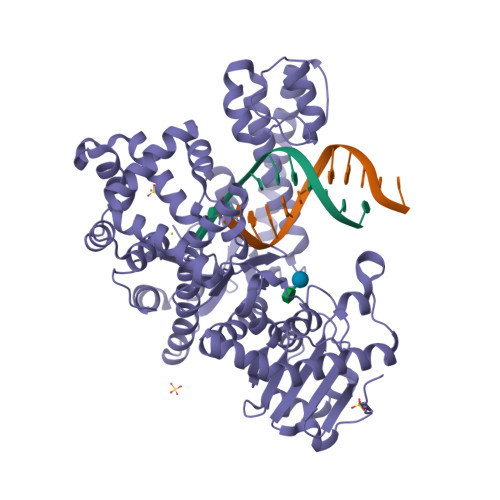
RCSB PDB - 1YFL: T4Dam in Complex with Sinefungin and 16-mer Oligonucleotide Showing Semi-specific and Specific Contact and Flipped Base
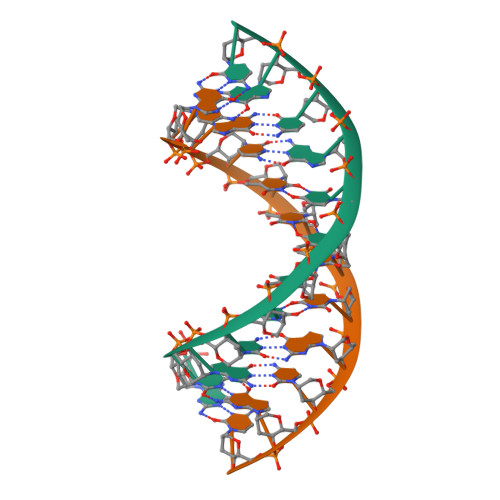
RCSB PDB - 1EC4: SOLUTION STRUCTURE OF A HEXITOL NUCLEIC ACID DUPLEX WITH FOUR CONSECUTIVE T:T BASE PAIRS
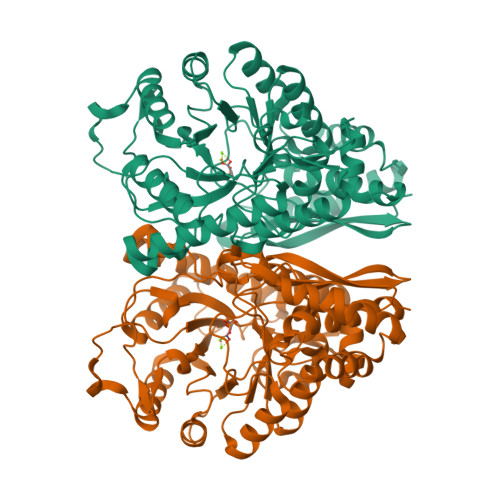
![RCSB PDB - 3O82: Structure of BasE N-terminal domain from Acinetobacter baumannii bound to 5'-O-[N-(2,3-dihydroxybenzoyl)sulfamoyl] adenosine RCSB PDB - 3O82: Structure of BasE N-terminal domain from Acinetobacter baumannii bound to 5'-O-[N-(2,3-dihydroxybenzoyl)sulfamoyl] adenosine](https://cdn.rcsb.org/images/structures/3o82_assembly-1.jpeg)
RCSB PDB - 3O82: Structure of BasE N-terminal domain from Acinetobacter baumannii bound to 5'-O-[N-(2,3-dihydroxybenzoyl)sulfamoyl] adenosine
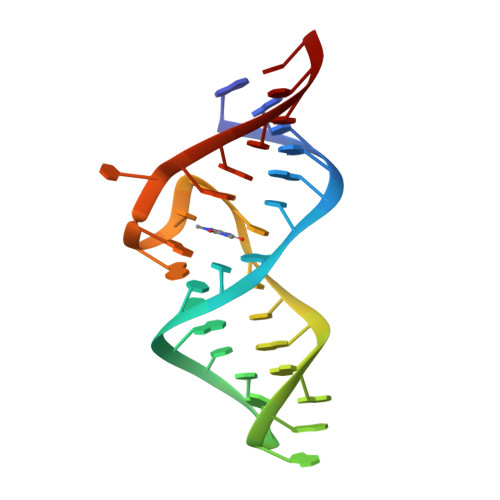
RCSB PDB - 1O15: THEOPHYLLINE-BINDING RNA IN COMPLEX WITH THEOPHYLLINE, NMR, REGULARIZED MEAN STRUCTURE, REFINEMENT WITH TORSION ANGLE AND BASE-BASE POSITIONAL DATABASE POTENTIALS AND DIPOLAR COUPLINGS
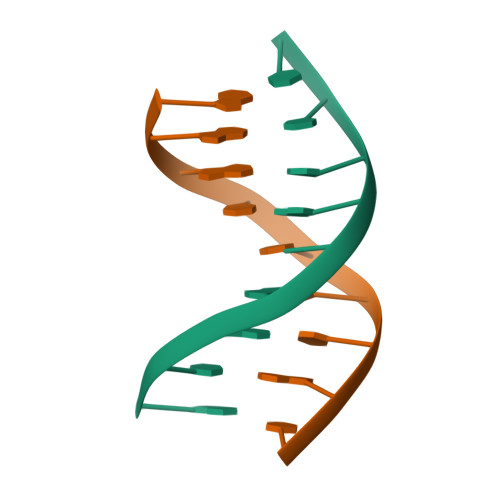
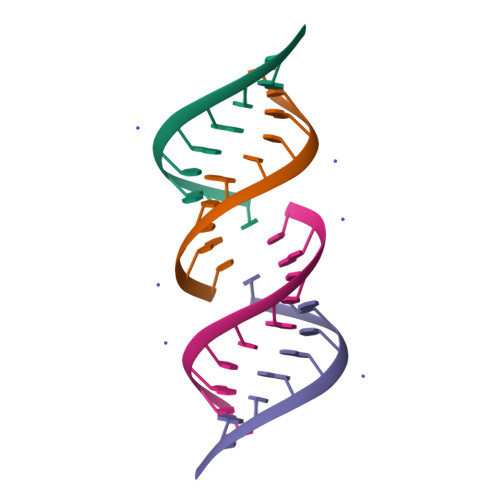
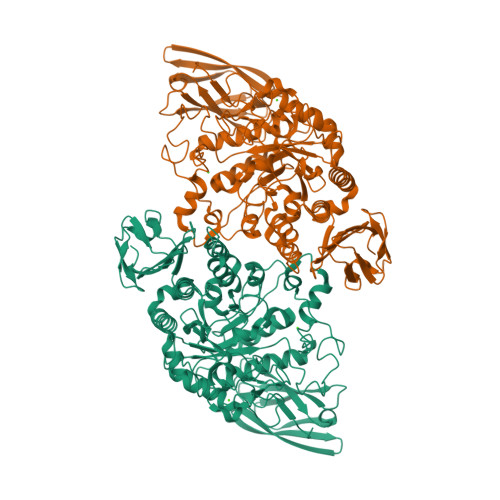



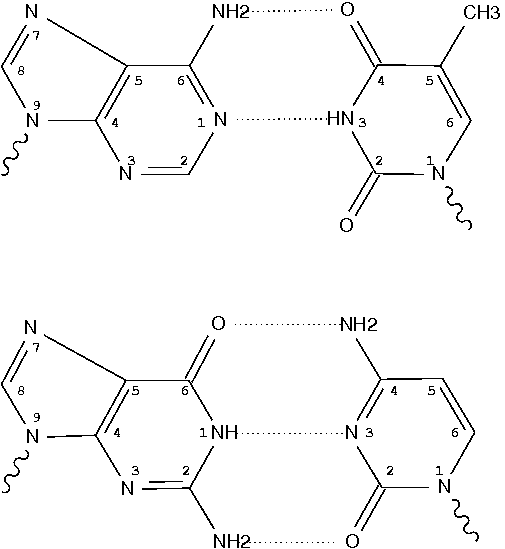








![RCSB PDB - 7SDF: [C:Ag+:S] Metal-mediated DNA base pair in a self-assembling rhombohedral lattice RCSB PDB - 7SDF: [C:Ag+:S] Metal-mediated DNA base pair in a self-assembling rhombohedral lattice](https://cdn.rcsb.org/images/structures/7sdf_assembly-1.jpeg)